There are large deviations from average values in some of the parameters. Are the values real? What is the best way to view them in structures if so ?
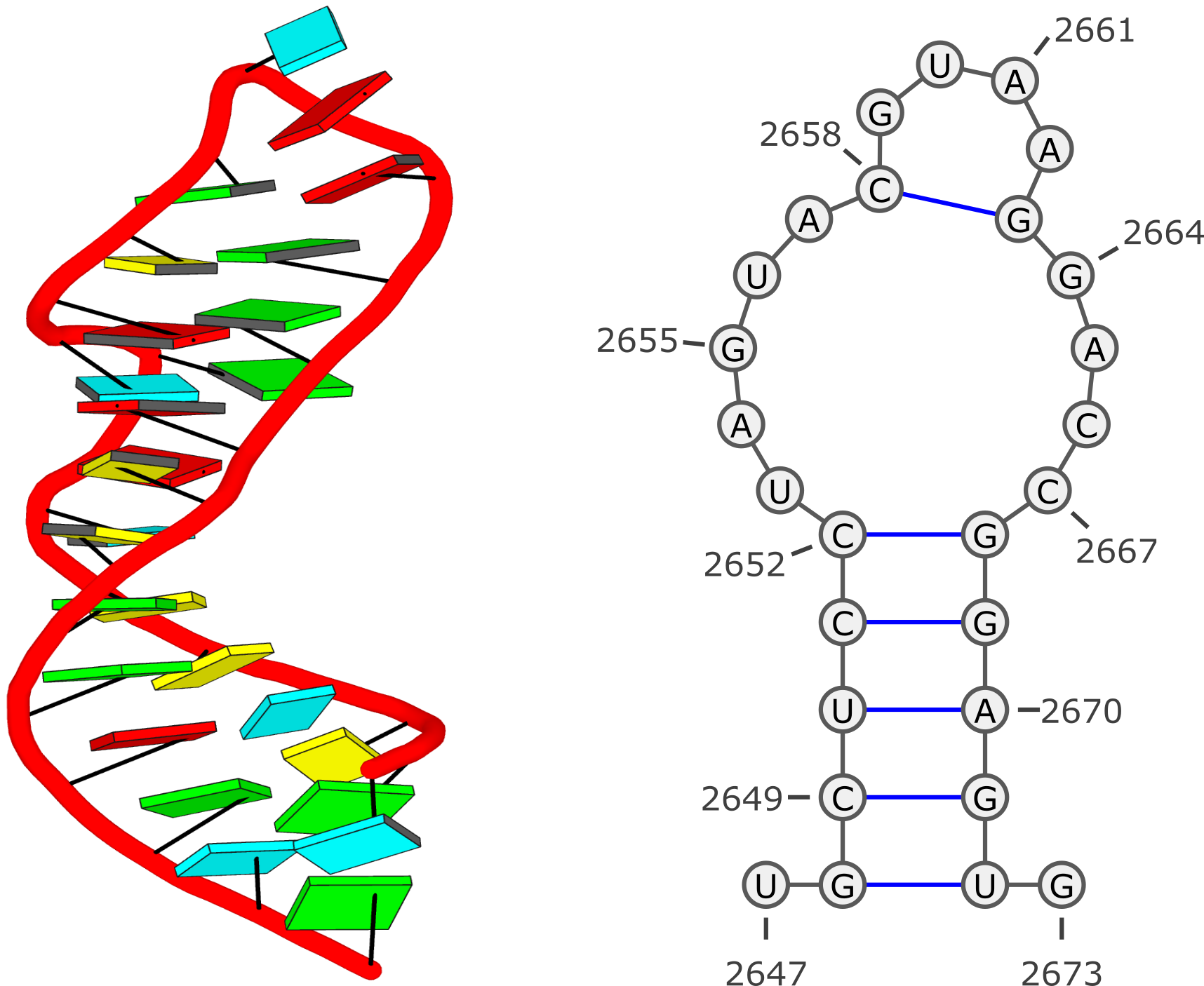
The short answer is yes, they are 'real', as would be expected from 3DNA. The large deviations of 3DNA parameters for 3sj2 (and many other PDB entries) are due to non-canonical base pairs (bp). In the case of 3sj2, they are the three G+G pairs.
3DNA adopts the
standard base reference frame, which is based on what would be a perfectly planar Watson-Crick (WC) bp geometry. Thus, by definition, WC pairs have the six bp parameters (Shear, Stretch, Stagger, Buckle, Propeller, Opening) close to zeros, with certain variations, mostly in Stagger, Buckle, and Propeller. Non-canonical bps can have numerous ways to deviate from a WC bp; however, whatever the case, they can be rigorously quantified by the six rigid-body bp parameters. See the
3DNA NAR03 paper for details.
A non-cannonical bp not only has a set of characteristic bp parameters (see below for G+G bps in 3sj2), it could also greatly effect the (middle) bp reference frame used to derive the bp step and helical parameters. That's why you see large deviations of those parameters from normal WC steps (see below).
bp Shear Stretch Stagger Buckle Propeller Opening
3 G-U -2.33 -0.47 0.13 2.30 -12.27 1.61
4 C-G 0.22 -0.06 -0.00 3.70 -18.31 2.91
5 C-G 0.22 -0.10 0.10 0.15 -4.59 -0.12
6 G+G -1.48 -3.61 -0.15 11.94 4.31 87.72
step Shift Slide Rise Tilt Roll Twist
4 CC/GG -0.61 -1.97 3.29 -1.87 9.30 30.50
5 CG/GG 0.04 -3.25 -1.34 -170.79 31.71 160.42
6 GG/CG -0.47 -3.69 -3.19 128.85 -110.61 97.02
3DNA derives a complete set of bp parameters to rigorously characterize the relative base geometry. For example, run the following 3DNA commands to see how '
analyze' and '
rebuild' complement each other to illustrate the point:
find_pair 3sj2.pdb stdout | analyze stdin
rebuild -atomic bp_step.par 3sj2-3dna.pdb
# superimpose '3sj2-3dna.pdb' onto '3sj2.pdb' using only base atoms, the rmsd would be ~0
find_pair 3sj2-3dna.pdb stdout | analyze stdin
# compare '3sj2-3dna.out' and '3sj2.out', the bp parameters are virtually identical
I see it as an advantage for 3DNA to report those 'weird' parameters, as it would (should) draw a user's attention. These bps and steps should be treated separately from the normal variations in structures (fragments) consisting of only WC bps. There could be
ad hoc ways to made 'weird' values look normal, but they lack rigor and consistency. Again, see the
3DNA NAR03 paper for more info.
Note also that for a relatively straight duplex structure like 3sj2, 3DNA also output a set of "Global parameters based on C1'-C1' vectors:" as shown below, which you may find useful:
disp.: displacement of the middle C1'-C1' point from the helix
angle: inclination between C1'-C1' vector and helix (subtracted from 90)
twist: helical twist angle between consecutive C1'-C1' vectors
rise: helical rise by projection of the vector connecting consecutive
C1'-C1' middle points onto the helical axis
bp disp. angle twist rise
1 G-C 8.82 12.28 28.16 3.19
2 G-C 8.09 11.46 32.47 2.62
3 G-U 6.75 11.40 28.94 2.40
4 C-G 7.06 8.02 31.16 2.87
5 C-G 7.45 8.60 33.16 2.88
6 G+G 7.07 7.95 29.95 2.72
7 G-C 7.20 8.14 30.17 3.24
8 C-G 6.86 9.94 34.81 2.83
9 G+G 6.13 10.17 28.18 2.62
10 G-C 6.56 6.75 33.11 2.91
11 C-G 6.71 6.30 30.30 2.70
12 G+G 6.05 9.99 33.37 2.80
13 G-C 6.48 10.27 29.88 3.13
14 G-C 6.82 7.48 27.46 2.97
15 U-G 7.18 5.34 38.30 2.79
16 C-G 7.16 10.13 26.27 2.90
17 C-G 7.17 12.43 --- --- HTH,
Xiang-Jun
 . Thus the discrepancy: you found that e-z can be negative in analyze -tor output, while I said that it's in range [0, 360] which should always be positive.
. Thus the discrepancy: you found that e-z can be negative in analyze -tor output, while I said that it's in range [0, 360] which should always be positive. . Thus the discrepancy: you found that e-z can be negative in analyze -tor output, while I said that it's in range [0, 360] which should always be positive.
. Thus the discrepancy: you found that e-z can be negative in analyze -tor output, while I said that it's in range [0, 360] which should always be positive.