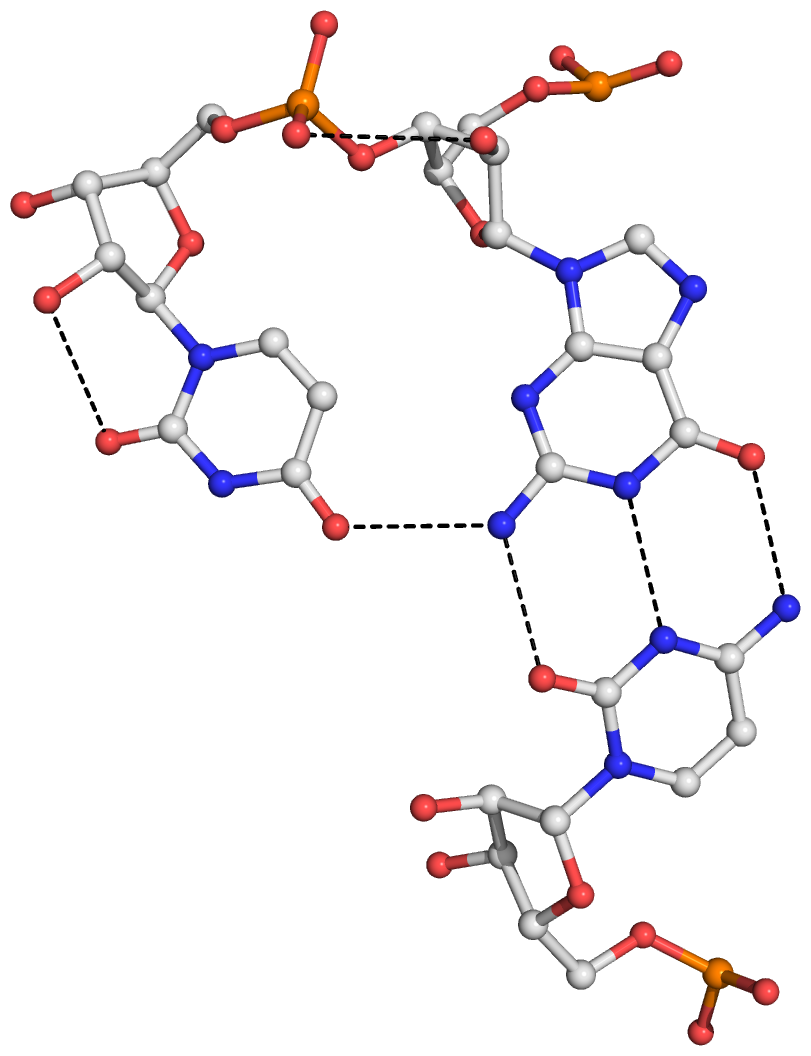
Figure S3: The four base triplets and associated hydrogen bonds (dashed lines) detected by DSSR in the viral tRNA mimic (PDB id: 4p5j) from turnip yellow mosaic virus. (A) GUC (G10,U11,C49) lies at the elbow of the L-shaped tertiary structure, with G10 and U11 forming a GpU dinucleotide platform. (B) CGU (C58,G72,U73) includes another GpU (G72 and U73) platform, with a single base-base hydrogen bond. (C) CGA (C59,G71,A75) forms a type I A-minor motif. (D) CGC (C61,G68,C77) contains a loop nucleotide (C61) from the hairpin-type pseudoknot. The last three triplets (B-D) are located at the 3'-end of the structure around the hairpin-type pseudoknot.
Starting from "
4p5j.pdb" downloaded from RCSB PDB, here is the complete script to get each of the four base-triplet images in png format.
x3dna-dssr -i=4p5j.pdb -o=4p5j.out --prefix=4p5j
ex_str -1 4p5j-multiplets.pdb 4p5j-m1.pdb
x3dna-dssr -i=4p5j-m1.pdb -o=4p5j-m1.pml --hbfile-pymol
pymol -qkc 4p5j-m1.pml
convert -trim +repage -border 10 -bordercolor white 4p5j-m1-pymol.png 4p5j-m1.png
ex_str -2 4p5j-multiplets.pdb 4p5j-m2.pdb
x3dna-dssr -i=4p5j-m2.pdb -o=4p5j-m2.pml --hbfile-pymol
pymol -qkc 4p5j-m2.pml
convert -trim +repage -border 10 -bordercolor white 4p5j-m2-pymol.png 4p5j-m2.png
ex_str -3 4p5j-multiplets.pdb 4p5j-m3.pdb
x3dna-dssr -i=4p5j-m3.pdb -o=4p5j-m3.pml --hbfile-pymol
pymol -qkc 4p5j-m3.pml
convert -trim +repage -border 10 -bordercolor white 4p5j-m3-pymol.png 4p5j-m3.png
ex_str -4 4p5j-multiplets.pdb 4p5j-m4.pdb
x3dna-dssr -i=4p5j-m4.pdb -o=4p5j-m4.pml --hbfile-pymol
pymol -qkc 4p5j-m4.pml
convert -trim +repage -border 10 -bordercolor white 4p5j-m4-pymol.png 4p5j-m4.png
Note:
- The --prefix option makes the auxiliary files having a specified prefix instead of the default "dssr". For example, "dssr-multiplets.pdb" becomes "4p5j-multiplets.pdb".
- The ex_str utility program is from the 3DNA distribution. It is used to extract a specific model from a MODEL/ENDMDL ensemble.
- The DSSR --hbfile-pymol option is used to generate a .pml file with all required settings for rendering in PyMOL.
- The convert program from ImageMagick is used here to trim extra white boundaries.
- The multiplet-png images (here four triplets) were combined using InkScape, and annotated, to get the final illustration.
- For completeness, here is the tarball file containing all the data files and the script ("tasks"): supp-fig3-4p5j-multiplets.tar.gz
Here is a sample image generated with the above script: